BESA Research
Best signal processing for your EEG and MEG data
- Product No: NS-1025
- Manufacturer: BESA
- EEG EP ERP MEG FFT
- Request Quote
Description
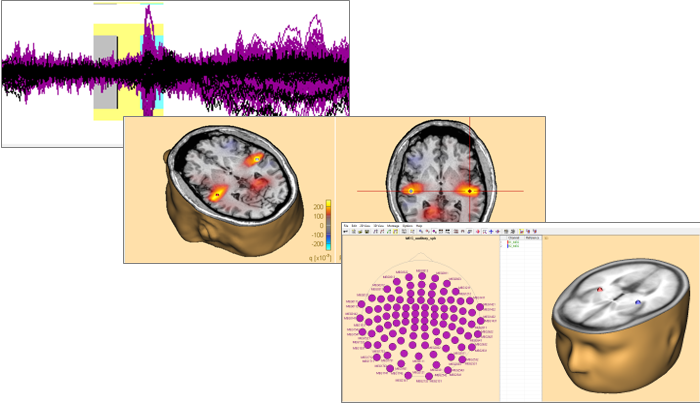
BESA is the most widely used software for source analysis and dipole localization in EEG and MEG research. BESA Research has been developed on the basis of 30 years experience in human brain research by Michael Scherg, University of Heidelberg, and Patrick Berg, University of Konstanz. BESA Research is a highly versatile and user-friendly Windows® program with optimized tools and scripts to preprocess raw or averaged data for source analysis. All important aspects of source analysis are displayed in one window for immediate selection of a wide range of tools. BESA Research provides fast and easy hypothesis testing, a variety of source analysis algorithms, integration with MRI and fMRI, standardized realistic head models (FEM) as well as the possibility to import individual head models (FEM) generated by BESA MRI 2.0. Source coherence, a unique feature for viewing brain activity BESA Research transforms surface signals into brain activity using source montages derived from multiple source models. This allows displaying ongoing EEG/MEG, single epochs, and averages with much higher spatial resolution. The Source Coherence Module provides an extremely fast and user-friendly implementation of time-frequency analysis based on complex demodulation. Users can create event-related time-frequency displays of power, amplitude, or event-related (de)synchronization and coherence for the current montage using brain sources or surface channels. Induced and evoked activities can be separated. Source coherence analysis reveals the functional connectivity between brain regions by reducing the volume conduction effects seen in surface coherence.
Data import and export
- Direct readers for most EEG and MEG data file formats
- Import of user-defined file formats using generic reader
- Data import / export to ASCII and binary files
- MATLAB interface for direct transfer of analysis results to MATLAB
Data processing
- Superior digital filtering: high, low, and narrow band pass, notch
- Interpolation from recorded to virtual and source channels
- Automated EOG and EKG artifact detection and correction
- Advanced user-defined instantaneous artifact correction
- Pattern detection and averaging by spatio-temporal correlation
Data review
- Easy and fast review of digital EEG and MEG data files
- Fast paging, tagging and selected viewing of epochs of interest
- DSA and event displays for quick jump to relevant pages
- Additional selected and virtual artifact channels (EOG etc.)
- Linear and non-linear correlation between scalp and source channels
- Spectral analysis: FFT, DSA, power and phase mapping
- Independent Component Analysis (ICA): Decomposition of EEG/MEG data into ICA components that can be used for artifact correction and as spatial sources in the source analysis window
Software Features
- Data review and processing
- Source montages and 3D whole-head mapping
- ERP analysis and averaging
- Source localization and source imaging
- Individual MRI and fMRI integration with BESA MRI and BrainVoyager
- Source coherence and time-frequency analysis
BESA Research 7.1 - New Features NEW
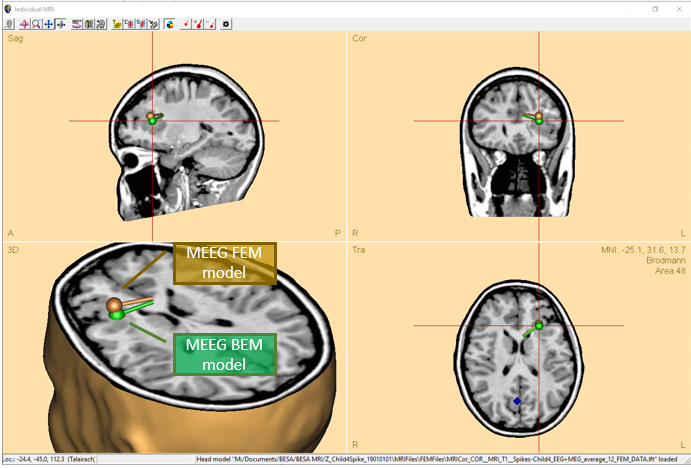
- Compare Boundary element method (BEM) and Finite element method (FEM) models on M/EEG data with just a couple of mouse clicks. These models are automatically loaded into the Source analysis module after calculating these in BESA MRI and can be exchanged in the head model section of the module.
- Combine MEG and EEG into a single data set for simultaneous and combined source modeling.
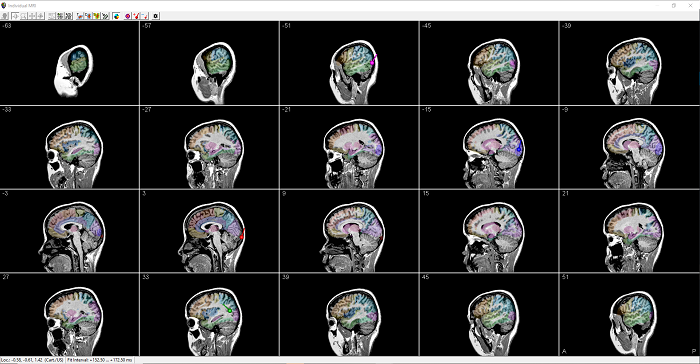
- For an easier overview of individual anatomies, MRI display now offers a multi-slice view in any orientation.
- Calculate confidence limits for discrete source solutions, and displayed these in co-registered MRI images.
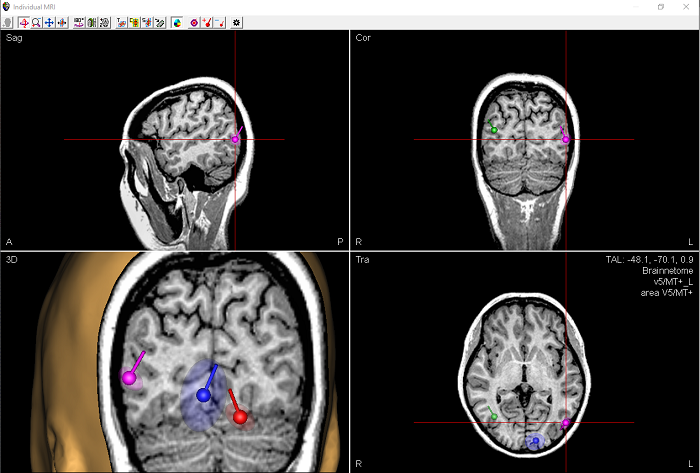
- Various improvements, enhancements, and speed optimizations for the Bayesian source imaging method SESAME were implemented. Furthermore, hyper-priors were introduced (see: https://arxiv.org/abs/2006.04141) and parallel computing was optimized.
- Utilise the full noise covariance matrix computed from individual trials in the computation of minimum norm estimates.
- Calculation of beamformer virtual sensor montages based on atlas regions is now supported.
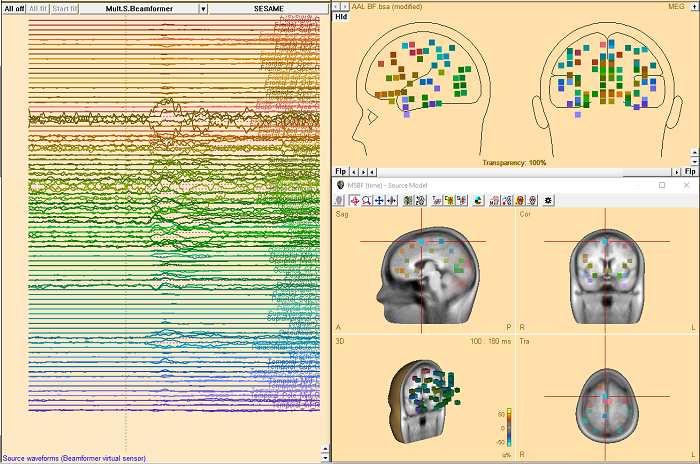
- Two new brain atlases added: Yeo7 and Yeo17 (https://doi.org/10.1152/jn.00338.2011).
- User Montreal Neurological Institute (MNI) coordinates now in the Source Analysis window.
- Automatic alerts for the baseline interval definition feature when it interferes with signals of interest.
- Color schemes for publication purposes are now available.
System Requirements
|
Operating system: |
Windows® 10, 8.1, 7 (64 bit, Touch not supported) |
|
Processor (CPU): |
Minimum: 2 GHz |
|
RAM: |
Minimum: 8 GB |
|
Screen resolution: |
Minimum: 1280x1024 pixels |
|
Graphics card: |
Supporting OpenGL 2.0 with 16 MB RAM or more |